"cmd.select" creates a named atom selection from a selection-expression. Command line instructions will be in "black" and clickable button instructions in blue. selName the name of the returned selection. list of up to 4-letter codes for atoms in proteins or nucleic acids PyMol> select Mycarbons, name ca+cb+cg+cd residue-name-list list of 3-letter codes for amino. residue-name-list list of 3-letter codes for amino acids PyMOL> select aas, resn ASP+GLU+ASN+GLN. atom-name-list list of up to 4-letter codes for atoms in proteins or nucleic acids PyMOL> select carbons, name CA+CB+CG+CD: resn: r. PyMOL> select polar, symbol O+N: name: n. The word "helix" is not defined in PyMol.! Restrict search to one atom per residue like this: iterate selected and name CA, ls.append ( (resi, resn)) Or, better yet, have the whole seqeunce in one line like this: iterate selected and name CA, print (resi, resn) Share. To see available options, you can type and then hit Tab, for. A typed PyMOL command always starts with a keyword that calls PyMOL to execute an action. Zoomout%again,sothat%youseethewholeprotein. Thereareseveral%different%waytopresent%youstructure,e.g.ascartoon, ribbons,%sticks%etc.%We%will%now%try%some%of%these.%% e.
#PYMOL TUTORIAL WIKI FULL#
Command line completion works inside of help, so if you don't remember the full keyword, type help, the first character or so of the keyword, and hit TAB. The script considers all molecules present in PyMOL individually. PyMOL> select active_water, ( (resi 38 and name ND2 and chain A) around 3.5) and (resn HOH) The above command select any water molecules that is/are around 3.5A of the ND2 atom of resi 38 in.
#PYMOL TUTORIAL WIKI WINDOWS#
In the command-line window (Depending on your PyMOL version, Windows labels this Tcl/Tk GUI or The PyMOL Molecular Graphics System), type the following commands: select AB, chain A+B hide all show cartoon, AB orient AB Note that in the panel at the top right, you now can operate on the subset AB using the buttons. Step 3: Use PyMol Useful for your images: This answer is not useful. User can also load the file through command line using the command load E.g., "load C: Downloads1UBQ.pdb" By default, the loaded PDB file structure will be shown with line representation in PyMol (Figure 2). Locate and Display the active site water We know that the amide group of Asn38 is h-bond to an active water. Regarding the L/H chain part, if they are effectively annotated as L and H, something like the following. To add a disulfide bond between amino acids 29 and 93: select 29 or 93 ssbonds 1.0. With Chime, you can use the select commands described here only on web pages which have a command line slot. But you can also create selections by the "select" command: "selection_expression" - An set of property name and value-list pairs joined by Boolean operators to identify a group of atoms in a molecule. The list of arguments for a command (the "usage") can be queried on the command line with a. If you then left-click on a neighboring residue, it too should be added to the selection: By using the script called "InterfaceResidues", you can select interface residues. Then select each of the residues using the mouse.
This work is licensed under a Creative Commons Attribution 4.0 International License: you must give appropriate credit, provide a link to the license, and indicate if changes were made.
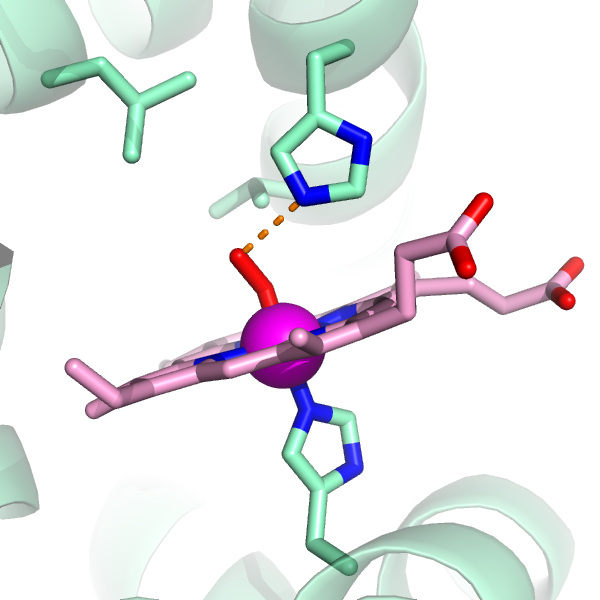
In the bottom right it says you are selecting "molecules" click on it and you will change to "chains". These commands allow the user to colour different parts of a protein and better visualise it in order to take better decisions on refining it. Can select them from the sequence Hide command can remove unwanted atoms. Luca Jovine has made available the program nuccyl, which allows PyMOL to display atomic models of nucleic acids in a highly simplified representation.Follow the above link for pictures, as well as a detailed tutorial on using the program. select creates a named selection from an atom selection. In PyMOL, users can easily add/create or delete chemical bonds. 1_visualize_model.py which visualizes the model and hightlight some regions and residues of interest by different colors. The command for selecting amino acids within x Å of a selection named selection1 is select NEARAA1, (byres expand x &selection1 !selection1). If you have set up PyMOL, you do not need to worry about finding and copying .Simply use the command "nuccyl" with the relevant arguments. The cartoon representation of the protein should now have turned white. needle the sequence of amino acids to find. You can also type help, for instance help color to get a man page for each command. That the active site has a pocket that accommodates an aromatic amino acid residue from the ligand.
